|
Here is a screenshot of the Analysis section: through this panel, the user can select the data file
and set the parameters that will be used in the analysis.
For example we suggest to use Legendre as basis function, maximum degree L of polynomial to be about half of the (distinct) time points available, to not provide smoothness information and hence set vi=0
To choose the parameter lambda of the truncated Poisson to match the expected prior degree, which is reasonable to lay between L/2 and L-1.
|



|
Bayesian Analysis for Time Series Microarray Experiments |





|
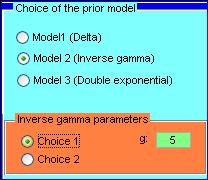
Here the user can select estimator for the global parameters (note that user specified choices are allowed) and chose the method for estimating the gene specific degree (two options are avaliable, MAP and Mean) And the range for maximizing the marginal probability distribution function of the data and hence estimate |
|
The user may control the Multiplicity error using the procedure proposed in Abramovich & Angelini, Bayesian MAP multiple testing procedures Sankhya (2006).
Binomial prior is usually found to be the most appropriate.
However, with the option Rank ordered is also possible to print out the ordered list of genes without automatically determine the cut-off of significance. The biologist can after decide to select only the top gene in the list without carrying out any formal test of significance. |
|
A user friendly software for Bayesian Analysis of Time Series Microarray Experiments. |

|
For a short description of the statistical model implemented in BATS see here |